How to Architect Medical Imaging AI Storage for Research
Medical imaging AI storage provides scalable, performant infrastructure for storing DICOM files, annotations, and model outputs for radiology and pathology AI applications. This guide explores architecture patterns for research and development workflows using AI agents.
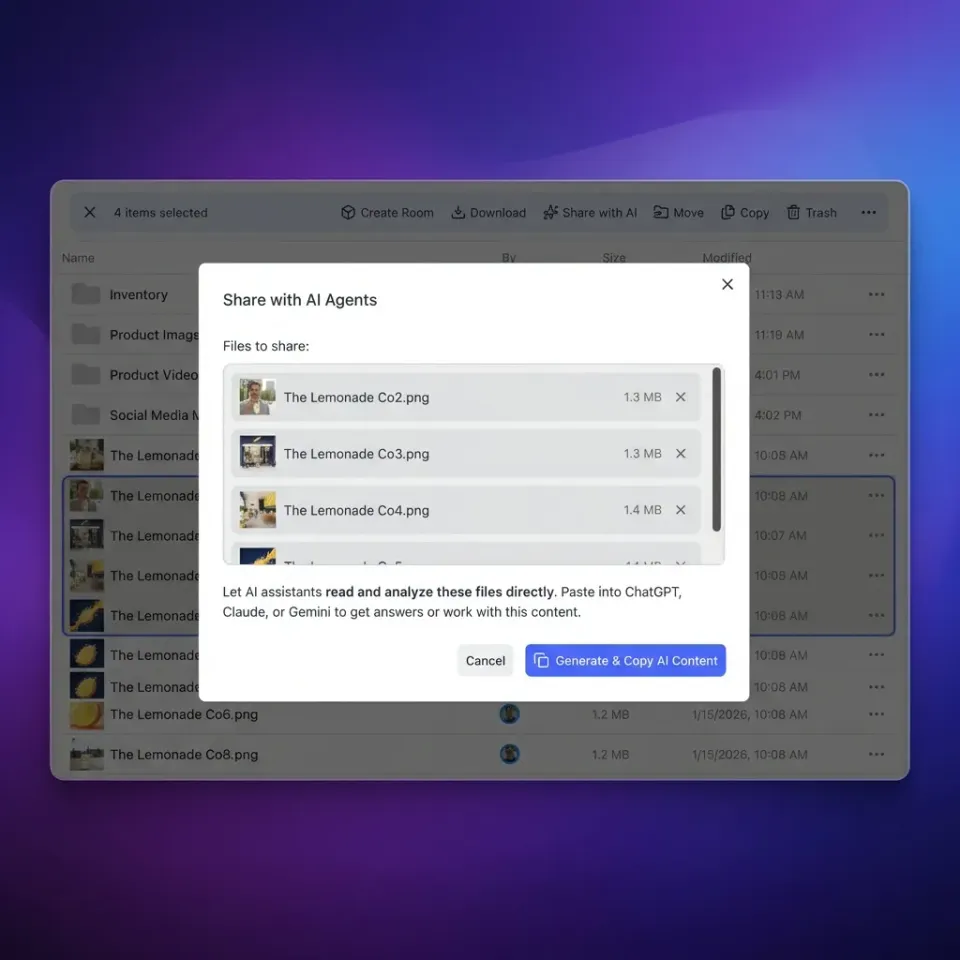
How to implement medical imaging AI storage reliably
Medical imaging data is growing fast. 3D mammography studies and pathology slides can generate large files. For AI development, this presents a dual challenge: storing petabytes of raw data while maintaining the high access speeds needed for model training.
High-resolution 3D volumes from MRI and CT scans require significant bandwidth. Digital pathology is even more demanding, with Whole Slide Imaging (WSI) generating files that can be 1-2 GB each. When training foundational models on large slide datasets, storage requirements reach hundreds of terabytes. This scale breaks traditional file server architectures, requiring object storage solutions that scale horizontally to handle the load.
Traditional PACS (Picture Archiving and Communication Systems) are built for clinical viewing, not the massive parallel ingestion required by machine learning pipelines. The medical imaging AI market is expected to reach $4.5 billion by 2027, led by research labs and tech companies that need more flexible storage solutions than hospital legacy systems can provide. The volume, velocity, and variety of data require a new storage strategy.
Helpful references: Fastio Workspaces, Fastio Collaboration, and Fastio AI.

What to check before scaling medical imaging AI storage
The most important architectural decision is separating clinical production environments from AI research and development sandboxes. Clinical environments require strict strict security requirements and integration with Electronic Health Records (EHR). Research environments, however, prioritize speed, API accessibility, and cost-efficiency for de-identified datasets.
The process of de-identification is not just about removing patient names. It requires scrubbing pixel data to remove burned-in annotations and cleaning complex DICOM header structures. Storage architecture plays a important role here, acting as the staging ground where "raw" anonymized data lands before being processed into "clean" training-ready formats like NIfTI or TFRecords. This separation creates a necessary "air gap" that protects patient privacy while speeding up research.
Fastio is designed specifically for the research and development side of this equation. While Fastio uses strong encryption and security measures, it is not strict security requirements compliant and should not be used for Protected Health Information (PHI). Instead, it acts as the logic layer for anonymized datasets where AI agents need unrestricted programmatic access to sort, label, and process files without the limits of clinical systems.

Accelerate Your AI Research for medical imaging AI storage
Get 50GB of free, high-performance storage for your de-identified datasets and AI agents.
Automating Data Curation with AI Agents
Modern storage architecture treats storage as an active participant in the workflow, not just a passive bucket. By using the Model Context Protocol (MCP), developers can deploy AI agents that directly interact with the file system to automate curation tasks.
For example, an AI agent connected via Fastio's MCP server can:
- Watch a "Drop Zone" folder for new de-identified upload batches
- Automatically validate file integrity and format (e.g., checking DICOM headers)
- Sort files into training, validation, and test directories based on metadata
- Trigger external processing pipelines via webhooks
Beyond simple sorting, agents can perform preliminary quality assurance. An agent could inspect every new image, check for sufficient contrast, resolution, or artifacting, and flag "blurry" scans for manual review. This "human-in-the-loop" workflow is key to creating high-quality ground truth datasets. This agent-native approach replaces simple Python scripts with intelligent, context-aware assistants that can handle exceptions and report status in natural language.
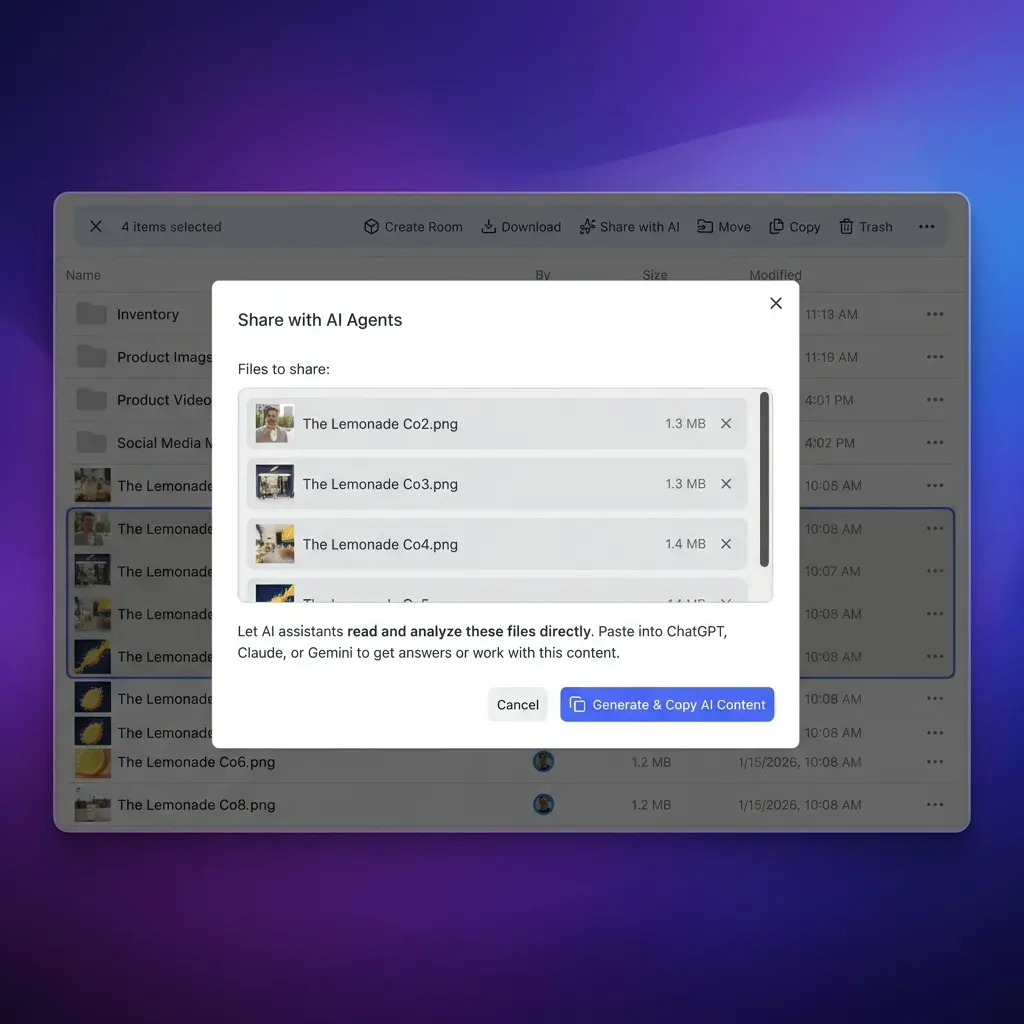
Hybrid Cloud Strategies for Imaging
Most research institutions use a hybrid approach where data originates on-premise at the modality (scanner) level but processing happens elsewhere. A common pattern is to use an on-premise gateway to buffer these studies, perform initial de-identification, and then asynchronously replicate them to cloud object storage.
This approach separates acquisition speed from internet bandwidth limits. Researchers can then mount the cloud bucket as a local drive for analysis or pull specific subsets into ephemeral high-performance cloud compute instances for model training sessions. This optimizes costs by not keeping high-performance GPUs idle while waiting for data transfer. It allows for a flexibility that purely on-premise or purely cloud-native solutions often struggle to match.
Performance and Data Gravity
"Data gravity" refers to the difficulty of moving large datasets. In medical imaging AI, moving terabytes of data to a compute cluster can take days. A modern architecture mitigates this through intelligent caching and edge delivery.
Latency slows down work. When a researcher opens a folder containing thousands of CT scans, waiting for the directory list to populate can break flow. Modern object storage with metadata acceleration ensures that file listing is instantaneous, feeling like a local SSD even when accessing data stored geographically distant.
Fastio's global edge network and HLS streaming capabilities allow distributed research teams to preview video-based medical data (like ultrasound or catheterization video) instantly without waiting for full downloads. For static images, the usage-based pricing model means research projects pay only for the storage and bandwidth they actually consume, rather than provisioning large storage systems that sit idle between training runs.
Security for Intellectual Property
While de-identified data removes patient privacy risks, the datasets themselves, and the AI models trained on them, represent massive intellectual property value. Securing this IP is important.
Data poisoning is a new threat where malicious actors introduce subtle artifacts into training data to corrupt model behavior. Immutable storage buckets with object locking (WORM - Write Once Read Many) provide a defense against this, ensuring that once a dataset is validated, it cannot be subtly altered without detection.
A strong storage architecture for AI IP includes:
- Granular Permissions: Role-based access control (RBAC) to ensure only authorized researchers access specific cohorts.
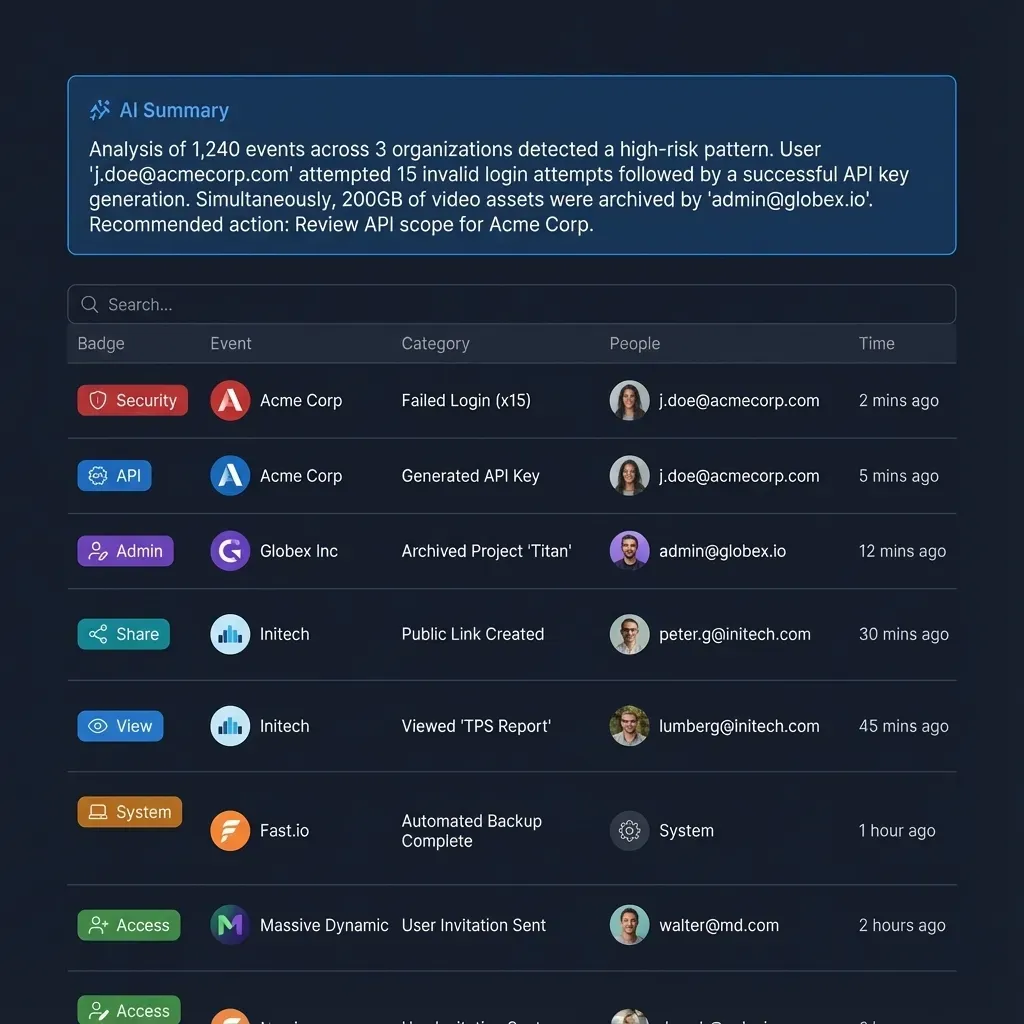
- Audit Logging: Comprehensive tracking of every file access, download, and modification to detect unauthorized theft.
- Encryption: Strong encryption at rest and in transit to protect proprietary models and training data.
Fastio includes these security features, ensuring that while the data is open to your agents, it remains closed to the world.
Frequently Asked Questions
Is Fastio strict security requirements compliant for medical images?
No, Fastio is not strict security requirements compliant. It is designed for high-performance storage of de-identified research data, public datasets, and non-clinical AI development. Do not store Protected Health Information (PHI) on Fastio.
Can I store DICOM files on Fastio?
Yes, Fastio supports any file type, including DICOM (.dcm) files. While we don't provide a built-in DICOM viewer, you can store, share, and programmatically access these files using standard APIs or AI agents.
How does AI storage differ from PACS?
PACS is optimized for clinical retrieval and human viewing in a hospital setting. AI storage focuses on high-throughput access for machine learning models, programmatic management, and massive scalability for training datasets.
What is the maximum file size for medical datasets?
Fastio supports massive files, making it ideal for large 3D imaging studies or whole-slide pathology images that can exceed several gigabytes. The platform is built to handle the heavy file sizes typical in medical research.
Can AI agents help organize medical datasets?
. Using the Fastio MCP server, you can build agents that automatically categorize, rename, and move files based on their content or metadata, reducing the manual labor involved in dataset preparation.
Related Resources

Accelerate Your AI Research for medical imaging AI storage
Get 50GB of free, high-performance storage for your de-identified datasets and AI agents.
